Example Molecules (click to use):
PFAS Groups Reference
You can download the current server PFAS groups JSON as a template. Any custom JSON you upload must follow the same structure (array of objects with keys like id, name, alias, main_group, smarts1, optional smarts2, componentSmarts, constraints, definition). The server sanitizes input to remove unknown keys and unsafe values.
About the PFAS Analyzer
The PFAS Analyzer is a web-based tool for identifying Per- and Polyfluoroalkyl Substances (PFAS) in molecular structures. It uses chemical structure patterns (SMARTS) and formula constraints to classify molecules into specific PFAS groups. This tool is designed for researchers, environmental scientists, and chemists working with fluorinated compounds.
The original work was developed in Python and should be preferred to JavaScript for production use. This web version is provided mainly for quick analyses and educational purposes for those who do not have access to Python or RDKit.
What are PFAS?
PFAS are a large group of synthetic chemicals that contain carbon-fluorine bonds. They are known as "forever chemicals" because they don't break down easily in the environment. PFAS are used in many industrial and consumer products, including non-stick cookware, water-repellent fabrics, firefighting foam, and food packaging. See this very brief introduction for more information.
- Identifies 28 OECD PFAS groups corresponding to the terminology in this report (figure 9) and 23 other more generic PFAS groups (see the tab on PFAS Groups).
- Supports SMILES and InChI input formats
- Batch processing via CSV/Excel import
- Export results with molecular structures
- Runs entirely in your browser (no data sent to servers)
Technology
This tool uses RDKit.js (a WebAssembly port of the RDKit cheminformatics library) for molecular structure processing, SMARTS pattern matching, and structure visualization. All analysis happens locally in your browser.
Citation & Credits
The tool was developed by Luc T. Miaz, part of the ZeroPM project (WP2) which received funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement No 101036756. This work was developed at the Department of Environmental Science at Stockholm University.



Based on the PFASgroups Python library. If you use this tool in your research, please cite the original publication.
🔬 Step-by-Step Algorithm Flow
The PFASGroupsJS algorithm detects PFAS functional groups by matching SMARTS patterns and analyzing fluorinated carbon components in molecular structures. The algorithm follows a multi-step process to identify which PFAS groups are present in a given molecule.
The algorithm identifies fluorinated components (chains of fluorinated carbons) and checks if functional groups (detected via SMARTS) are connected to these components. This approach ensures that only truly fluorinated functional groups are detected.
Algorithm Steps
1Input & Preprocessing
- Parse SMILES or InChI string into molecular structure
- Add explicit hydrogens to molecule (critical for formula calculation)
- Calculate molecular formula and parse into element dictionary
2Molecule Sanitization
- Attempt to sanitize the molecule (validate valences)
- If sanitization fails, fragment the molecule to fix valence errors
- Continue with valid fragments
3PFAS Group Iteration
For each PFAS group definition in the database:
- Formula Constraint Check: Verify molecular formula satisfies group constraints (relational, minimum/maximum counts, element restrictions)
- SMARTS Pattern Matching: Check if functional groups are present using substructure searches
- Component Overlap: Check if matched SMARTS atoms are within distance=2 of fluorinated components
For PFAS groups to be detected, the SMARTS pattern must match an atom that is part of the fluorinated component or satisfies the component criteria. This means:
- The matched atom should be a carbon that is per- or polyfluorinated, OR
- The matched atom must be within
max_dist_from_CFdistance from the nearest C-F carbon - Example: ✅
[C$(C(=O)O)]where the matched C is bonded to CF₂ groups - Example: ❌ Functional group atoms completely isolated from the fluorinated chain won't be detected
Telomer groups are exceptions: they use linker_smarts to explicitly
define non-fluorinated linkers (e.g., CH₂ chains) between the perfluorinated chain and
the functional group.
4Result Recording
- Record all matching PFAS groups
- Count number of matches for each group
- Store component information with graph metrics
Formula Constraint Types
| Type | Description | Example |
|---|---|---|
| rel | Relational constraint between elements | C = F/2 + 0.5 (carbon equals half of fluorines plus 0.5) |
| gte | Greater than or equal (minimum) | F ≥ 1 (at least one fluorine atom) |
| lte | Less than or equal (maximum) | H ≤ 2 (at most two hydrogens) |
| eq | Equal to (exact count) | O = 2 (exactly two oxygen atoms) |
| only | Only these elements allowed | only: [C, F, H] (no other elements permitted) |
🔬 PFAS Groups Taxonomy
Interactive visualization of OECD and Generic PFAS group classifications. Click on any group to see SMARTS patterns, constraints, and parent-child relationships.
Fluorinated Chain Pathways
For some PFAS groups, the analyzer looks for fluorinated chains connecting two functional groups.
The pathway patterns defined in fpaths.json specify which atoms are considered part
of a valid fluorinated path.
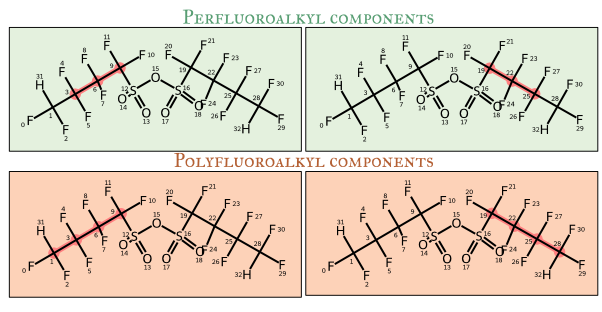
Pathway Types
1. Perfluoroalkyl Paths
Fully fluorinated carbon chains where every carbon has only C-F or C-C bonds (no C-H, C-Cl, etc.). Used for identifying PFCAs, PFSAs, and related compounds.
SMARTS Pattern:
[#6$([#6H0X4]([F,$(CF),$(CCF),$([#8]CF)])([!#17,!#35,!#54])([!#17,!#35,!#54])[C,F]),#8$([#8]CF)]
2. Polyfluoroalkyl Paths
Partially fluorinated carbon chains that may contain C-H bonds in addition to C-F bonds. Used for identifying PolyFCAs, PolyFSAs, and fluorotelomer compounds.
SMARTS Pattern:
[#6$([CX4]([#9,$(C[#9]),$(CC[#9]),$([#8]CF)])[C,F,#1,#17,#35,#53]),#8$([#8]CF)]
Path Finding Algorithm
- Identify all atoms matching
smarts1(e.g., carboxylic acid group) - Identify all atoms matching
smarts2(e.g., another functional group or chain end) - Find shortest path between each pair of matched atoms
- Check if all atoms in the path match the pathway SMARTS pattern
- Record valid paths as fluorinated chains
Example: PFOA (Perfluorooctanoic acid)
Structure: CF₃-CF₂-CF₂-CF₂-CF₂-CF₂-CF₂-COOH
Functional group (smarts1): -COOH (carboxylic acid)
Chain end (smarts2): -CF₃ (trifluoromethyl)
Path: All carbon atoms between -COOH and -CF₃
Pattern match: All carbons are fully fluorinated
Result: Matches "Perfluoroalkyl" pathway → Classified as PFCA
⭐ Molecule Prioritization
The prioritization tool ranks analyzed molecules by similarity to reference compounds or by intrinsic fluorination characteristics. This is useful for screening large datasets, regulatory watchlist generation, and environmental monitoring applications.
Prioritization Modes
Ranks molecules by distributional similarity to a reference list using KL divergence on PFAS group fingerprints.
- Use case: Find molecules similar to known persistent PFAS (e.g., PFOA, PFOS)
- Method: Compares PFAS group composition patterns
- Lower scores = more similar to reference
Formula: KL(reference || molecule) = Σ pref(g) × log(pref(g) / pmol(g))
Where pref and pmol are the normalized PFAS group frequencies.
Ranks molecules by fluorination characteristics using a weighted scoring function.
- Use case: Identify highly fluorinated or long-chain PFAS
- Method: Combines total fluorine content and chain length metrics
- Higher scores = more priority
Formula: Score = a × Σ(component sizes) + b × percentile(component sizes, p)
Parameters:
• a: Weight for total fluorination (sum of all CFn units)
• b: Weight for longest/largest chains (percentile-based)
• p: Percentile to focus on (e.g., 90 = top 10% of chains)
Parameter Guidelines
| Objective | Recommended Settings |
|---|---|
| Environmental persistence | a=1.0, b=5.0, p=90 (favor long chains) |
| Bioaccumulation potential | a=2.0, b=3.0, p=75 (balanced) |
| High fluorine content | a=5.0, b=1.0, p=50 (favor total F) |
| Longest single chain | a=0.0, b=10.0, p=100 (only max) |
Workflow
- Analyze molecules: Run normal PFAS group analysis first
- Enable prioritization: Check "Enable prioritization" in the prioritization section
- Choose mode:
- For similarity: Check "Use reference list" and enter reference SMILES
- For intrinsic: Uncheck reference mode and adjust a, b, p parameters
- Apply: Click "Apply Prioritization" to rank results
- Export: Priority scores are included in CSV/Excel exports
Example Applications
- Analyze your dataset of molecules
- Enable prioritization with reference list
- Enter PFOA and PFOS SMILES as references:
C(=O)(C(C(C(C(C(C(C(F)(F)F)(F)F)(F)F)(F)F)(F)F)(F)F)(F)F)O C(C(C(C(C(C(C(C(F)(F)S(=O)(=O)O)(F)F)(F)F)(F)F)(F)F)(F)F)(F)F)(F)(F)F
- Apply prioritization - molecules with similar group patterns rank higher
- Analyze your dataset
- Enable prioritization without reference list
- Set parameters: a=1.0, b=5.0, p=90
- Apply - molecules with longest chains appear first
Interpreting Scores
- Reference mode (KL divergence): Lower scores indicate greater similarity. Score of 0 = identical composition.
- Intrinsic mode: Higher scores indicate higher priority based on your chosen weighting.
- Equal scores: Molecules with identical fluorination profiles receive the same priority.
Python Integration
The full Python package provides additional prioritization capabilities:
- Statistical analysis of priority distributions
- Batch processing of large databases
- Integration with SQL databases
- Advanced filtering and clustering
See the Python documentation for details.
Usage Guide
Single Molecule Analysis
- Select input format (SMILES or InChI)
- Enter molecular structure in the text area
- Click "Analyze" or press Ctrl+Enter
- View results with structure visualization and detected groups
Batch Processing
- Prepare a CSV or Excel file with a column containing SMILES or InChI strings
- Click "📁 Click to import CSV or Excel file"
- Select your file (column auto-detection happens automatically)
- Click "Analyze" to process all molecules
- Export results to CSV or Excel with optional structure images
Export Options
- Include all columns: Creates separate Yes/No columns for each PFAS group type
Example Input Formats
SMILES Examples:
# PFOA (Perfluorooctanoic acid)
C(=O)(C(C(C(C(C(C(C(F)(F)F)(F)F)(F)F)(F)F)(F)F)(F)F)(F)F)O
# PFOS (Perfluorooctanesulfonic acid)
C(C(C(C(C(C(C(C(F)(F)S(=O)(=O)O)(F)F)(F)F)(F)F)(F)F)(F)F)(F)F)(F)(F)F
# 6:2 FTOH
C(C(C(C(C(C(F)(F)F)(F)F)(F)F)(F)F)(F)F)(F)(F)CCO
CSV File Structure:
Name,SMILES,CAS,Category
PFOA,C(=O)(C(C(...))O,335-67-1,PFCA
PFOS,C(C(C(...))F,1763-23-1,PFSA
GenX,C(C(C(OC(...))O,13252-13-6,PFECA
Keyboard Shortcuts
- Ctrl+Enter: Analyze molecules (when text area is focused)
- Use the example molecules to test the analyzer
- Check browser console (F12) for detailed debugging information
Troubleshooting
| Issue | Solution |
|---|---|
| Invalid molecule structure | Check SMILES/InChI format; try copying from a chemical database |
| No PFAS groups detected | Molecule may not contain fluorine or doesn't match PFAS patterns |
| Import file not working | Ensure file has header row; check that molecule column contains valid structures |
| Slow performance | For very large datasets, consider splitting into smaller batches |
| Freeze | Keep the tab on focus during execution of the allgorithm. If needed check browser console (F12) for detailed debugging information |
🔧 Key Functions
matchAllSmarts(mol, smartsDict)
Matches all SMARTS patterns from a group's definition and checks minimum count requirements.
/** * Match all SMARTS patterns from a group's smarts dictionary * Returns Set of all atoms that match any SMARTS pattern */ function matchAllSmarts(mol, smartsDict) { if (!smartsDict || typeof smartsDict !== 'object') { return null; } const allMatchedAtoms = new Set(); for (const [smartsPattern, minCount] of Object.entries(smartsDict)) { let pattern = RDKit.get_qmol(smartsPattern); const matchesJson = mol.get_substruct_matches(pattern); pattern.delete(); // Check if we have at least minCount matches if (matches.length < minCount) { return null; // Not enough matches } // Add all matched atoms to set for (const match of matches) { for (const atomIdx of match.atoms) { allMatchedAtoms.add(atomIdx); } } } return allMatchedAtoms.size > 0 ? allMatchedAtoms : null; }
connectedComponents(graph, subset)
Finds connected components in a molecular graph using BFS (Breadth-First Search).
/** * Find connected components in a molecular graph using BFS * Takes a graph and a subset of node indices * Returns array of Sets (each Set is a component) */ function connectedComponents(graph, subset) { const subsetSet = new Set(subset); const visited = new Set(); const components = []; // BFS to find each connected component for (const startNode of subsetSet) { if (visited.has(startNode)) continue; const component = new Set(); const queue = [startNode]; while (queue.length > 0) { const node = queue.shift(); const neighbors = graph.edges.get(node) || []; for (const edge of neighbors) { const neighborIdx = edge.neighbor; // Fixed: was edge.to if (subsetSet.has(neighborIdx) && !visited.has(neighborIdx)) { visited.add(neighborIdx); component.add(neighborIdx); queue.push(neighborIdx); } } } components.push(component); } return components; }
isWithinDistance (Distance-2 Connectivity)
Checks if a matched atom is within a specified distance of a fluorinated component using BFS.
/** * Check if atom is within distance N of a component using BFS * This allows functional groups 2 bonds away from fluorinated chains * to be correctly associated with those chains */ function isWithinDistance(startAtom, component, graph, maxDistance) { if (!graph || !graph.edges) return false; const visited = new Set([startAtom]); let queue = [{ atom: startAtom, distance: 0 }]; while (queue.length > 0) { const { atom, distance } = queue.shift(); if (component.has(atom)) { return true; // Found connection! } if (distance < maxDistance) { const neighbors = graph.edges.get(atom) || []; for (const edge of neighbors) { if (!visited.has(edge.to)) { visited.add(edge.to); queue.push({ atom: edge.to, distance: distance + 1 }); } } } } return false; }
⚖️ Python vs JavaScript Implementation
Comparison of key differences between the Python reference implementation and this JavaScript port.
| Aspect | Python (Reference) | JavaScript (Current) | Impact |
|---|---|---|---|
| Hydrogen Atoms | Always adds explicit H with Chem.AddHs() | Attempts add_hs() but may fail silently | HIGH - Affects SMARTS matching |
| Component Definition | Uses NetworkX connected_components() | Custom BFS implementation | LOW - Logic equivalent |
| Adjacency Check | Distance-based BFS with maxDistance=2 | Distance-based BFS with maxDistance=2 | LOW - Now equivalent (fixed!) |
| SMARTS Matching | mol.GetSubstructMatches() | mol.get_substruct_matches() | LOW - API equivalent |
| Graph Building | RDKit mol_to_nx() | Parse molblock manually | MEDIUM - May miss some edges |
| Formula Extraction | Chem.rdMolDescriptors.CalcMolFormula() | Parse molblock or provided formula | MEDIUM - May miss implicit H |
| Component Merging | Universal merging across pathways | Universal merging across pathways | LOW - Now equivalent (fixed!) |
- Fix #1: Universal Component Merging
Now merges all fluorinated atoms across pathways before computing connected components, preventing fragmentation in branched molecules. - Fix #2: Distance-2 Connectivity
Expanded adjacency check from distance=1 to distance=2, catching functional groups on carbons adjacent to fluorinated components. - Fix #3: Graph Edge Property
Fixed connectedComponents to use edge.neighbor instead of edge.to, matching the graph structure from molToGraph.
- Explicit hydrogen handling may not be 100% reliable
- Formula extraction from molblock may miss some implicit hydrogens
- Manual graph building could miss exotic bond types
- Linker validation not supported: The JavaScript version cannot validate specific atoms between fluorinated components and functional groups (linker_smarts feature)
- Subset of PFAS groups: This JavaScript version includes core PFAS groups but not all specialized groups (e.g., fluorotelomer variants groups 60-73)
- For production use and complete capabilities, use the Python implementation which includes all 73+ PFAS groups and full linker validation support
🔍 JS vs Python: Behaviour Summary
This tab summarises how closely this JavaScript port follows the Python PFASGroups reference implementation, which changes were made during the porting/parity work, and in which situations the results can still differ.
- PFAS group definitions (IDs 1–73 with
compute=true) are shared with the Python package via the same JSON configuration. - The component graph logic (connected components, distance-based extension, adjacency tests) mirrors the Python NetworkX-based solver using a custom BFS implementation.
- Validated on full OECD dataset: 1559 out of 1994 valid compounds (78.2%) have identical PFAS group assignments between Python and JavaScript.
- The remaining 435 molecules (21.8%) differ primarily in edge-case functional groups or complex hydrogen handling scenarios, not in core PFAS scaffold detection.
Changes implemented in this JavaScript port
- Shared PFAS group definitions: JavaScript reads the same PFAS group JSON definitions as Python for IDs 1–73, including SMARTS patterns, formula constraints and component types.
- Groups 18 (Perfluoroalkenes) and 19 (Hydrofluoroolefins): Their SMARTS patterns, distance limits (
max_dist_from_CF) and linker settings were aligned exactly with the Python configuration to avoid inconsistent classification of double-bonded systems. - Universal component merging: All fluorinated atoms from different pathway definitions are merged before computing connected components, matching the Python behaviour on branched or overlapping chains.
- Distance-2 connectivity: Functional groups up to two bonds away from a fluorinated component are associated with that component, reproducing the Python adjacency criterion.
- Graph edge handling: The connected‐component routine was corrected to follow the same neighbour semantics as the Python NetworkX graph, preventing artificial fragmentation.
- Perfluoro vs polyfluoro overlap handling: When perfluoroalkyl and polyfluoroalkyl components describe the same atom set, only the polyfluoro component is kept, mirroring Python’s preference for the more general class.
Where results can still differ from Python
- Hydrogen and formula handling: Python uses Chem.AddHs and CalcMolFormula, while JavaScript reconstructs formulas from molblocks and RDKit.js. In edge cases this can change whether certain H-dependent patterns (e.g. some acids, amides or chlorides) are recognised.
- Generic functional groups: The JavaScript version may sometimes report additional generic groups (e.g. alcohols, esters, sulfonic acids or side-chain aromatics) attached to a fluorinated skeleton even when the Python implementation is more conservative.
- Specific functional groups: For a few molecules the Python implementation reports groups such as hydrofluoroolefins (19), perfluoro dicarboxylic acids (3), certain amides (35) or halides (43) that the JavaScript implementation currently misses or classifies differently.
- Linker patterns: Python can enforce detailed linker SMARTS between fluorinated components and functional groups. The JavaScript port approximates this behaviour with distance-based checks but does not fully support all linker validation logic.
- Subset of PFAS groups: This web tool focuses on core PFAS groups. Some specialised groups present in Python (for example certain fluorotelomer variants in the 60–73 range) are not evaluated here.
Practical guidance
- Use this JavaScript analyzer for quick screening, exploration and educational purposes, especially when Python/RDKit are not available.
- For regulatory, inventory or production workflows where exact reproducibility is critical, prefer the Python PFASGroups package as the reference implementation.
- When results differ slightly between Python and JavaScript, differences are most likely in a small number of generic functional groups rather than in the presence/absence of a PFAS scaffold.